Synthesizes Short Segments Of Rna
As discussed in Chapter 3, DNA replication is a semiconservative procedure in which each parental strand serves as a template for the synthesis of a new complementary girl strand. The fundamental enzyme involved is Deoxyribonucleic acid polymerase, which catalyzes the joining of deoxyribonucleoside 5′-triphosphates (dNTPs) to form the growing Deoxyribonucleic acid chain. All the same, DNA replication is much more than complex than a single enzymatic reaction. Other proteins are involved, and proofreading mechanisms are required to ensure that the accuracy of replication is compatible with the low frequency of errors that is needed for prison cell reproduction. Additional proteins and specific DNA sequences are also needed both to initiate replication and to re-create the ends of eukaryotic chromosomes.
Deoxyribonucleic acid Polymerases
DNA polymerase was first identified in lysates of East. coli by Arthur Kornberg in 1956. The ability of this enzyme to accurately copy a DNA template provided a biochemical basis for the manner of Deoxyribonucleic acid replication that was initially proposed by Watson and Crick, so its isolation represented a landmark discovery in molecular biological science. Ironically, nevertheless, this beginning DNA polymerase to be identified (now called Dna polymerase I) is not the major enzyme responsible for E. coli DNA replication. Instead, it is now articulate that both prokaryotic and eukaryotic cells contain several different Dna polymerases that play distinct roles in the replication and repair of DNA.
The multiplicity of DNA polymerases was first revealed by the isolation of a mutant strain of E. coli that was deficient in polymerase I (Figure 5.1). Cultures of East. coli were treated with a chemic (a mutagen) that induces a high frequency of mutations, and individual bacterial colonies were isolated and screened to place a mutant strain defective polymerase I. Analysis of a few chiliad colonies led to the isolation of the desired mutant, which was almost totally defective in polymerase I activity. Surprisingly, the mutant bacteria grew normally, leading to the determination that polymerase I is not required for Deoxyribonucleic acid replication. On the other mitt, the mutant bacteria were extremely sensitive to agents that damage Dna (east.g., ultraviolet light), suggesting that polymerase I is involved primarily in the repair of Dna damage rather than in DNA replication per se.

Effigy 5.1
Isolation of a mutant scarce in polymerase I. A culture of East. coli was treated with a chemic mutagen, and individual bacterial colonies were isolated by growth on semisolid medium. Several grand colonies were then cultured and screened to identify (more...)
The conclusion that polymerase I is not required for replication unsaid that Eastward. coli must contain other DNA polymerases, and subsequent experiments led to the identification of two such enzymes, now called DNA polymerases Two and Three. The potential roles of these enzymes were investigated by the isolation of appropriate mutants. Strains of E. coli with mutations in polymerase 2 were found to grow and otherwise conduct normally, so the role of this enzyme in the prison cell is unknown. Temperature-sensitive polymerase III mutants, however, were unable to replicate their DNA at high temperature, and subsequent studies have confirmed that polymerase Three is the major replicative enzyme in E. coli.
Information technology is at present known that, in add-on to polymerase III, polymerase I is also required for replication of Eastward. coli DNA. The original polymerase I mutant was not completely defective in that enzyme, and later experiments showed that the residual polymerase I activeness in this strain plays a key role in the replication process. The replication of E. coli Deoxyribonucleic acid thus involves two singled-out DNA polymerases, the specific roles of which are discussed below.
Eukaryotic cells incorporate 5 DNA polymerases: α, β, γ, δ, and ε. Polymerase γ is located in mitochondria and is responsible for replication of mitochondrial Dna. The other 4 enzymes are located in the nucleus and are therefore candidates for involvement in nuclear DNA replication. Polymerases α, δ, and ε are nigh active in dividing cells, suggesting that they function in replication. In contrast, polymerase β is agile in nondividing and dividing cells, suggesting that it may function primarily in the repair of Dna damage.
Two types of experiments accept provided further prove addressing the roles of polymerases α, δ, and ε in Dna replication. Get-go, replication of the DNAs of some animal viruses, such as SV40, tin be studied in cell-costless extracts. The ability to study replication in vitro has immune direct identification of the enzymes involved, and assay of such jail cell-free systems has shown that polymerases α and δ are required for SV40 Deoxyribonucleic acid replication. 2nd, polymerases α, δ, and ε are found in yeasts also as in mammalian cells, enabling the use of the powerful approaches of yeast genetics (see Chapter 3) to test their biological roles directly. Such studies indicate that yeast mutants lacking any of these three Dna polymerases are unable to proliferate, implying a disquisitional role for polymerase ε likewise as for α and δ. Notwithstanding, further studies have shown that the essential function of polymerase ε in yeast does not require its activeness every bit a replicative DNA polymerase. Thus, polymerases α and δ appear to be sufficient for Dna replication both in jail cell-gratis systems and in yeast, so the role of polymerase ε remains to be established.
All known DNA polymerases share 2 primal properties that deport critical implications for DNA replication (Figure 5.ii). First, all polymerases synthesize Deoxyribonucleic acid only in the v′ to 3′ direction, calculation a dNTP to the 3′ hydroxyl grouping of a growing concatenation. Second, DNA polymerases tin can add a new deoxyribonucleotide only to a preformed primer strand that is hydrogen-bonded to the template; they are not able to initiate DNA synthesis de novo past catalyzing the polymerization of gratis dNTPs. In this respect, Deoxyribonucleic acid polymerases differ from RNA polymerases, which can initiate the synthesis of a new strand of RNA in the absenteeism of a primer. As discussed later in this affiliate, these properties of DNA polymerases announced critical for maintaining the high fidelity of DNA replication that is required for cell reproduction.
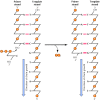
Figure v.2
The reaction catalyzed by Dna polymerase. All known Deoxyribonucleic acid polymerases add a deoxyribonucleoside 5′-triphosphate to the iii′ hydroxyl group of a growing Dna chain (the primer strand).
The Replication Fork
DNA molecules in the process of replication were kickoff analyzed by John Cairns in experiments in which E. coli were grown in the presence of radioactive thymidine, which allowed subsequent visualization of newly replicated Dna by autoradiography (Figure five.iii). In some cases, complete circular molecules in the process of replicating could be observed. These Dna molecules contained ii replication forks, representing the regions of active Deoxyribonucleic acid synthesis. At each fork the parental strands of DNA separated and two new girl strands were synthesized.

Figure v.3
Replication of E. coli Dna. (A) An autoradiograph showing leaner that were grown in [threeH]thymidine for ii generations to label the DNA, which was and so extracted and visualized by exposure to photographic pic. (B) This schematic illustrates the two (more...)
The synthesis of new Deoxyribonucleic acid strands complementary to both strands of the parental molecule posed an important problem to understanding the biochemistry of DNA replication. Since the ii strands of double-helical DNA run in reverse (antiparallel) directions, continuous synthesis of ii new strands at the replication fork would crave that 1 strand be synthesized in the 5′ to iii′ direction while the other is synthesized in the reverse (3′ to 5′) management. But Dna polymerase catalyzes the polymerization of dNTPs merely in the 5′ to 3′ direction. How, then, can the other progeny strand of DNA be synthesized?
This enigma was resolved by experiments showing that only one strand of DNA is synthesized in a continuous style in the direction of overall DNA replication; the other is formed from small, discontinuous pieces of DNA that are synthesized backward with respect to the direction of movement of the replication fork (Figure 5.4). These pocket-sized pieces of newly synthesized Deoxyribonucleic acid (chosen Okazaki fragments after their discoverer) are joined past the action of DNA ligase, forming an intact new Deoxyribonucleic acid strand. The continuously synthesized strand is called the leading strand, since its elongation in the direction of replication fork movement exposes the template used for the synthesis of Okazaki fragments (the lagging strand).

Figure five.4
Synthesis of leading and lagging strands of Dna. The leading strand is synthesized continuously in the management of replication fork movement. The lagging strand is synthesized in small pieces (Okazaki fragments) backward from the overall direction of (more than...)
Although the discovery of discontinuous synthesis of the lagging strand provided a mechanism for the elongation of both strands of DNA at the replication fork, it raised another question: Since Dna polymerase requires a primer and cannot initiate synthesis de novo, how is the synthesis of Okazaki fragments initiated? The answer is that brusque fragments of RNA serve as primers for DNA replication (Figure 5.5). In contrast to DNA synthesis, the synthesis of RNA can initiate de novo, and an enzyme called primase synthesizes short fragments of RNA (eastward.g., iii to ten nucleotides long) complementary to the lagging strand template at the replication fork. Okazaki fragments are then synthesized via extension of these RNA primers past Deoxyribonucleic acid polymerase. An important consequence of such RNA priming is that newly synthesized Okazaki fragments contain an RNA-DNA joint, the discovery of which provided disquisitional show for the office of RNA primers in DNA replication.

Figure v.5
Initiation of Okazaki fragments with RNA primers. Short fragments of RNA serve as primers that can be extended by Dna polymerase.
To form a continuous lagging strand of DNA, the RNA primers must somewhen be removed from the Okazaki fragments and replaced with Dna. In E. coli, RNA primers are removed by the combined activity of RNase H, an enzyme that degrades the RNA strand of RNA-DNA hybrids, and polymerase I. This is the aspect of Due east. coli DNA replication in which polymerase I plays a critical role. In addition to its DNA polymerase action, polymerase I acts as an exonuclease that can hydrolyze DNA (or RNA) in either the 3′ to five′ or 5′ to 3′ management. The action of polymerase I every bit a 5′ to 3′ exonuclease removes ribonucleotides from the 5′ ends of Okazaki fragments, allowing them to be replaced with deoxyribonucleotides to yield fragments consisting entirely of DNA (Effigy 5.6). In eukaryotic cells, other exonucleases accept the identify of E. coli polymerase I in removing primers, and the gaps between Okazaki fragments are filled by the action of polymerase δ. As in prokaryotes, these DNA fragments can and so be joined by DNA ligase.
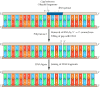
Figure 5.half dozen
Removal of RNA primers and joining of Okazaki fragments. Because of its 5′ to iii′ exonuclease activeness, Deoxyribonucleic acid polymerase I removes RNA primers and fills the gaps between Okazaki fragments with Dna. The resultant DNA fragments tin then exist (more...)
The different Deoxyribonucleic acid polymerases thus play distinct roles at the replication fork (Figure 5.7). In prokaryotic cells, polymerase III is the major replicative polymerase, functioning in the synthesis both of the leading strand of DNA and of Okazaki fragments by the extension of RNA primers. Polymerase I then removes RNA primers and fills the gaps between Okazaki fragments. In eukaryotic cells, still, two Deoxyribonucleic acid polymerases are required to do what in East. coli is accomplished by polymerase III alone. Polymerase α is institute in a complex with primase, and it appears to function in conjunction with primase to synthesize short RNA-Deoxyribonucleic acid fragments during lagging strand synthesis. Polymerase δ can so synthesize both the leading and lagging strands, acting to extend the RNA-Dna primers initially synthesized by the polymerase α-primase complex. In add-on, polymerase δ can take the place of E. coli polymerase I in filling the gaps betwixt Okazaki fragments following primer removal.

Figure 5.7
Roles of DNA polymerases in E. coli and mammalian cells. The leading strand is synthesized by polymerase III (politico 3) in Eastward. coli and past polymerase δ (politician δ) in mammalian cells. In Due east. coli, lagging strand synthesis is initiated by primase, (more than...)
Not only polymerases and primase but also a number of other proteins act at the replication fork. These additional proteins have been identified both by the analysis of East. coli mutants defective in DNA replication and by the purification of the mammalian proteins required for in vitro replication of SV40 DNA. 1 class of proteins required for replication binds to DNA polymerases, increasing the activity of the polymerases and causing them to remain bound to the template Deoxyribonucleic acid so that they continue synthesis of a new Deoxyribonucleic acid strand. Both Due east. coli polymerase III and eukaryotic polymerase δ are associated with ii types of accompaniment proteins (sliding-clamp proteins and clench-loading proteins) that load the polymerase onto the primer and maintain its stable clan with the template (Figure five.eight). The clamp-loading proteins (called the γ complex in E. coli and replication cistron C [RFC] in eukaryotes) specifically recognize and bind Deoxyribonucleic acid at the junction between the primer and template. The sliding-clench proteins (β protein in E. coli and proliferating jail cell nuclear antigen [PCNA] in eukaryotes) bind adjacent to the clamp-loading proteins, forming a ring around the template DNA. The clench proteins so load the Deoxyribonucleic acid polymerase onto Dna at the primer-template junction. The ring formed by the sliding clench maintains the association of the polymerase with its template as replication gain, allowing the uninterrupted synthesis of many thousands of nucleotides of Deoxyribonucleic acid.

Figure 5.8
Polymerase accessory proteins. (A) The clamp-loading protein (RFC in mammalian cells) binds DNA at the junction between primer and template. The sliding-clamp poly peptide (PCNA in mammalian cells) binds next to the RFC, and DNA polymerase then binds to (more than...)
Other proteins unwind the template DNA and stabilize unmarried-stranded regions (Figure v.ix). Helicases are enzymes that catalyze the unwinding of parental Dna, coupled to the hydrolysis of ATP, ahead of the replication fork. Single-stranded Deoxyribonucleic acid-binding proteins (due east.g., eukaryotic replication factor A [RFA]) so stabilize the unwound template Dna, keeping it in an extended unmarried-stranded country and so that it can be copied by the polymerase.
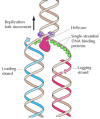
Figure 5.9
Action of helicases and single-stranded DNA-bounden proteins. Helicases unwind the two strands of parental DNA ahead of the replication fork. The unwound Deoxyribonucleic acid strands are then stabilized past single-stranded Deoxyribonucleic acid-binding proteins so that they can serve as (more...)
Every bit the strands of parental DNA unwind, the DNA ahead of the replication fork is forced to rotate. Unchecked, this rotation would cause circular DNA molecules (such every bit SV40 DNA or the E. coli chromosome) to go twisted around themselves, eventually blocking replication (Figure 5.x). This trouble is solved by topoisomerases, enzymes that catalyze the reversible breakage and rejoining of DNA strands. There are two types of these enzymes: Type I topoisomerases break simply ane strand of Deoxyribonucleic acid; type II topoisomerases introduce simultaneous breaks in both strands. The breaks introduced by type I and type 2 topoisomerases serve as "swivels" that allow the two strands of template Dna to rotate freely around each other so that replication can proceed without twisting the DNA ahead of the fork (encounter Figure 5.10). Although eukaryotic chromosomes are composed of linear rather than circular DNA molecules, their replication also requires topoisomerases; otherwise, the complete chromosomes would have to rotate continually during DNA synthesis.

Figure v.10
Action of topoisomerases during DNA replication. (A) Equally the 2 strands of template Deoxyribonucleic acid unwind, the DNA ahead of the replication fork is forced to rotate in the contrary management, causing circular molecules to become twisted around themselves. (B) This (more...)
Type Two topoisomerase is needed not only to unwind DNA simply likewise to unravel newly replicated circular DNA molecules that become interwined with each other. In eukaryotic cells, topoisomerase Ii appears to be involved in mitotic chromosome condensation. In improver, studies of yeast mutants, besides as experiments in Drosophila and mammalian cells, indicate that topoisomerase 2 is required for the separation of daughter chromatids at mitosis, suggesting that it functions to untangle newly replicated loops of DNA in the chromosomes of eukaryotes.
The enzymes involved in DNA replication act in a coordinated fashion to synthesize both leading and lagging strands of DNA simultaneously at the replication fork (Figure 5.11). This job is accomplished by the germination of dimers of the replicative DNA polymerases (polymerase III in E. coli or polymerase δ in eukaryotes), each with its appropriate accessory proteins. 1 molecule of polymerase and then acts in synthesis of the leading strand while the other acts in synthesis of the lagging strand. The lagging strand template is thought to form a loop at the replication fork so that the polymerase subunit engaged in lagging strand synthesis moves in the same overall management as the other subunit, which is synthesizing the leading strand.
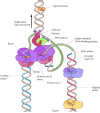
Figure 5.eleven
Model of the Eastward. coli replication fork. Helicase, primase, and two molecules of Deoxyribonucleic acid polymerase III acquit out coordinated synthesis of both the leading and lagging strands of Dna. The lagging strand template is folded so that the polymerase responsible (more than...)
The Fidelity of Replication
The accuracy of DNA replication is disquisitional to cell reproduction, and estimates of mutation rates for a diverseness of genes signal that the frequency of errors during replication corresponds to just one wrong base per 109 to 1010 nucleotides incorporated. This error frequency is much lower than would be predicted simply on the basis of complementary base pairing. In detail, the standard configurations of nucleic acrid bases are in equilibrium with rare culling conformations (tautomeric forms) that hydrogen-bond with the wrong partner (due east.yard., Chiliad with T) with a frequency of nearly one incorrect base per 104 (Figure 5.12). The much higher caste of fidelity actually achieved results largely from the activities of Deoxyribonucleic acid polymerase.

Figure 5.12
Mismatching between rare configurations of nucleic acid bases. In its normal configuration, guanine (K) specifically forms complementary hydrogen bonds (dashed lines) with cytosine (C). All the same, G occasionally assumes a rare configuration (tautomeric (more...)
One mechanism by which Deoxyribonucleic acid polymerase increases the fidelity of replication is by helping to select the right base for insertion into newly synthesized Deoxyribonucleic acid. The polymerase does not simply catalyze incorporation of whatever nucleotide is hydrogen-bonded to the template strand. Instead, it actively discriminates against incorporation of a mismatched base, presumably by adapting to the conformation of a right base pair. The molecular mechanisms responsible for the ability of Deoxyribonucleic acid polymerases to select against incorrect bases are not yet entirely understood, but this selectivity appears to increase the accuracy of replication about a hundredfold, reducing the expected error frequency from 10-iv to approximately 10-6.
The other major machinery responsible for the accuracy of Dna replication is the proofreading activity of DNA polymerase. Equally already noted, East. coli polymerase I has iii′ to 5′ equally well every bit 5′ to 3′ exonuclease activeness. The 5′ to 3′ exonuclease operates in the direction of DNA synthesis and helps remove RNA primers from Okazaki fragments. The 3′ to 5′ exonuclease operates in the reverse direction of DNA synthesis, and participates in proofreading newly synthesized DNA (Figure five.13). Proofreading is effective considering Dna polymerase requires a primer and is non able to initiate synthesis de novo. Primers that are hydrogen-bonded to the template are preferentially used, so when an incorrect base is incorporated, it is likely to be removed past the 3′ to v′ exonuclease activity rather than being used to keep synthesis. Such three′ to 5′ exonuclease activities are besides associated with Eastward. coli polymerase III and eukaryotic polymerases δ and ε. The three′ to 5′ exonucleases of these polymerases selectively excise mismatched bases that have been incorporated at the end of a growing Dna chain, thereby increasing the accuracy of replication by a hundred- to a thousandfold.
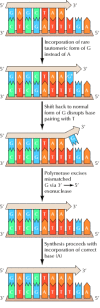
Figure 5.13
Proofreading by DNA polymerase. A rare tautomeric grade of 1000 (Chiliad*) is incorporated in place of A as a result of mispairing with T on the template strand. The subsequent shift of G dorsum to its normal form disrupts this base pairing, then the iii′ terminal (more...)
The importance of proofreading may explicate the fact that Deoxyribonucleic acid polymerases crave primers and catalyze the growth of DNA strands but in the 5′ to 3′ direction. When Deoxyribonucleic acid is synthesized in the five′ to 3′ direction, the energy required for polymerization is derived from hydrolysis of the 5′ triphosphate group of a free dNTP as it is added to the three′ hydroxyl group of the growing concatenation (see Figure v.2). If Deoxyribonucleic acid were to be extended in the three′ to 5′ direction, the energy of polymerization would instead take to exist derived from hydrolysis of the v′ triphosphate grouping of the terminal nucleotide already incorporated into Deoxyribonucleic acid. This would eliminate the possibility of proofreading, considering removal of a mismatched terminal nucleotide would likewise remove the v′ triphosphate group needed as an energy source for further chain elongation. Thus, although the ability of DNA polymerase to extend a primer only in the 5′ to 3′ direction appears to make replication a complicated process, it is necessary for ensuring authentic duplication of the genetic fabric.
Combined with the power to discriminate against the insertion of mismatched bases, the proofreading activity of DNA polymerases is sufficient to reduce the error frequency of replication to about i mismatched base per ten9. Additional mechanisms (discussed in the section "Deoxyribonucleic acid Repair") act to remove mismatched bases that have been incorporated into newly synthesized DNA, further ensuring right replication of the genetic information.
Origins and the Initiation of Replication
The replication of both prokaryotic and eukaryotic DNAs starts at a unique sequence called the origin of replication, which serves as a specific binding site for proteins that initiate the replication procedure. The get-go origin to be divers was that of E. coli, in which genetic analysis indicated that replication e'er begins at a unique site on the bacterial chromosome. The E. coli origin has since been studied in detail and establish to consist of 245 base pairs of DNA, elements of which serve as bounden sites for proteins required to initiate DNA replication (Figure v.14). The fundamental stride is the bounden of an initiator poly peptide to specific Deoxyribonucleic acid sequences within the origin. The initiator protein begins to unwind the origin Deoxyribonucleic acid and recruits the other proteins involved in Deoxyribonucleic acid synthesis. Helicase and single-stranded Dna-binding proteins and so act to go on unwinding and exposing the template Deoxyribonucleic acid, and primase initiates the synthesis of leading strands. 2 replication forks are formed and move in reverse directions along the round E. coli chromosome.

Figure five.fourteen
Origin of replication in East. coli. Replication initiates at a unique site on the E. coli chromosome, designated the origin (ori). The start event is the bounden of an initiator protein to ori DNA, which leads to fractional unwinding of the template. The Deoxyribonucleic acid (more...)
The origins of replication of several fauna viruses, such as SV40, have been studied as models for the initiation of Deoxyribonucleic acid synthesis in eukaryotes. SV40 has a single origin of replication (consisting of 64 base of operations pairs) that functions both in infected cells and in cell-free systems. Replication is initiated past a virus-encoded protein (called T antigen) that binds to the origin and besides acts as a helicase. A single-stranded DNA-binding protein is required to stabilize the unwound template, and the Dna polymerase α-primase complex then initiates DNA synthesis.
Although single origins are sufficient to direct the replication of bacte-rial and viral genomes, multiple origins are needed to replicate the much larger genomes of eukaryotic cells within a reasonable menstruation of fourth dimension. For instance, the entire genome of E. coli (four × 106 base pairs) is replicated from a unmarried origin in approximately xxx minutes. If mammalian genomes (3 × 10nine base pairs) were replicated from a single origin at the aforementioned charge per unit, DNA replication would require about iii weeks (30,000 minutes). The problem is further exacerbated by the fact that the rate of Deoxyribonucleic acid replication in mammalian cells is actually about tenfold lower than in Eastward. coli, perchance equally a event of the packaging of eukaryotic Deoxyribonucleic acid in chromatin. Nonetheless, the genomes of mammalian cells are typically replicated within a few hours, necessitating the use of thousands of replication origins.
The presence of multiple replication origins in eukaryotic cells was first demonstrated by the exposure of cultured mammalian cells to radioactive thymidine for different time intervals, followed by autoradiography to detect newly synthesized Deoxyribonucleic acid. The results of such studies indicated that Dna synthesis is initiated at multiple sites, from which it and so proceeds in both directions along the chromosome (Figure 5.15). The replication origins in mammalian cells are spaced at intervals of approximately l to 300 kb; thus the man genome has virtually 30,000 origins of replication. The genomes of simpler eukaryotes also take multiple origins; for instance, replication in yeasts initiates at origins separated by intervals of approximately xl kb.
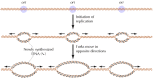
Figure v.15
Replication origins in eukaryotic chromosomes. Replication initiates at multiple origins (ori), each of which produces two replication forks.
The origins of replication of eukaryotic chromosomes have been studied best in yeasts, in which they have been identified as sequences that can back up the replication of plasmids in transformed cells (Effigy five.xvi). This has provided a functional assay for these sequences, and several such elements (called autonomously replicating sequences, or ARSs) take been isolated. Their role equally origins of replication has been verified by straight biochemical analysis, not but in plasmids just also in yeast chromosomal DNA.
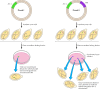
Effigy 5.16
Identification of origins of replication in yeast. Both plasmids I and II contain a selectable mark gene (LEU ii) that allows transformed cells to grow on medium lacking leucine. Only plasmid Ii, even so, contains an origin of replication (ARS). The (more...)
Functional ARS elements span about 100 base of operations pairs, including an 11-base-pair core sequence common to many dissimilar ARSs (Effigy 5.17). This cadre sequence is essential for ARS function and has been found to be the binding site of a protein circuitous (chosen the origin replication circuitous, or ORC) that is required for initiation of DNA replication at yeast origins. The ORC complex appears to recruit other proteins (including Dna helicases) to the origin, leading to the initiation of replication. The mechanism of initiation of DNA replication in yeasts thus appears similar to that in prokaryotes and eukaryotic viruses; that is, an initiator poly peptide specifically binds to origin sequences.

Figure v.17
A yeast ARS element. The element contains an 11-base-pair ARS consensus sequence (ACS), which is the specific bounden site of the origin replication complex (ORC). 3 boosted elements (B1, B2, and B3) are individually non essential but together (more than...)
In contrast to the well-defined ARS elements in yeasts, much less is known most the nature of replication origins in more complex eukaryotes. Nevertheless, recent experiments accept shown that specific origin sequences are required for initiation of Dna replication in mammalian cells. In addition, proteins related to the yeast ORC proteins have been identified in a variety of eukaryotes, including Drosophila, C. elegans, Arabidopsis, and humans, and shown to be essential for DNA replication. It thus appears likely that the basic mechanism used to initiate Deoxyribonucleic acid replication is conserved in eukaryotic cells.
Telomeres and Telomerase: Replicating the Ends of Chromosomes
Because Dna polymerases extend primers but in the 5′ to 3′ direction, they are unable to copy the extreme five′ ends of linear Dna molecules. Consequently, special mechanisms are required to replicate the terminal sequences of the linear chromosomes of eukaryotic cells. These sequences (telomeres) consist of tandem repeats of elementary-sequence DNA (meet Affiliate 4). They are replicated by the action of a unique enzyme called telomerase, which is able to maintain telomeres by catalyzing their synthesis in the absence of a Dna template.
Telomerase is a reverse transcriptase, ane of a class of DNA polymerases, first discovered in retroviruses (see Chapter 3), that synthesize DNA from an RNA template. Importantly, telomerase carries its own template RNA, which is complementary to the telomere repeat sequences, equally part of the enzyme complex. The employ of this RNA equally a template allows telomerase to generate multiple copies of the telomeric repeat sequences, thereby maintaining telomeres in the absence of a conventional Dna template to direct their synthesis.
The mechanism of telomerase action was initially elucidated in 1985 past Carol Greider and Elizabeth Blackburn during studies of the protozoan Tetrahymena (Figure 5.18). The Tetrahymena telomerase is complexed to a 159-nucleotide-long RNA that includes the sequence 3′-AACCCCAAC-5′. This sequence is complementary to the Tetrahymena telomeric repeat (5′-TTGGGG-3′) and serves equally the template for the synthesis of telomeric DNA. The use of this RNA as a template allows telomerase to extend the 3′ end of chromosomal Deoxyribonucleic acid by one echo unit of measurement beyond its original length. The complementary strand tin can and then exist synthesized by the polymerase α-primase complex using conventional RNA priming. Removal of the RNA primer leaves an overhanging 3′ end of chromosomal DNA, which can form loops at the ends of eukaryotic chromosomes (run into Figure 4.19).
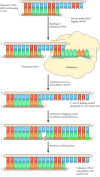
Figure v.xviii
Action of telomerase. Telomeric Dna is a simple repeat sequence with an overhanging 3′ end on the newly synthesized leading strand. Telomerase carries its own RNA molecule, which is complementary to telomeric Deoxyribonucleic acid, as part of the enzyme circuitous. (more...)
Telomerase has been identified in a diversity of eukaryotes, and genes encoding telomerase RNAs have been cloned from Tetrahymena, yeasts, mice, and humans. In each example, the telomerase RNA contains sequences complementary to the telomeric repeat sequence of that organism (come across Table four.3). Moreover, the introduction of mutant telomerase RNA genes into yeasts has been shown to result in corresponding alterations of the chromosomal telomeric echo sequences, directly demonstrating the role of telomerase in maintaining the termini of eukaryotic chromosomes.

Synthesizes Short Segments Of Rna,
Source: https://www.ncbi.nlm.nih.gov/books/NBK9940/
Posted by: vazquezanythat.blogspot.com


0 Response to "Synthesizes Short Segments Of Rna"
Post a Comment